| treatment | tree1 | tree2 | tree3 | tree4 | tree5 |
|---|---|---|---|---|---|
| oil | 4 | 29 | 14 | 14 | 7 |
| spores | -4 | 29 | 4 | -2 | 11 |
| water | -9 | 18 | 10 | 9 | -6 |
Analysis of the Complete Block Design
2024-10-09
Announcements
- Grades posted for HW4
- HW5 posted on Friday
- MP1 pre-approval due Friday 11:55p
- Office hours (Bass 412)
- Tuesday 10:30a - 11:30a
- Friday office hours cancelled (again)
- Thursday 4p-5p or by appointment
MP1 Pre-Approval
Your completed Qualtrics experiments are due for Pre-Approval by Fri Oct 11 11:59p.
- Export your survey as a Word doc. Include the survey flow and any skip logic you used.
- Upload the Word doc to Moodle.
I should be able to see any images you used or links to media. I’ll be paying particular attention to your consent form. It’s not unusual for me to ask for revisions before you launch your study.
Next steps:
I will give you approval or revisions via Slack direct messages. Watch out for those!
Once you have completed any revisions and receive approval from me you can complete the following checklist and then share your anonymous survey link.
- Proof read for errors.
- Test (preview) your survey multiple times, creating fake data.
- Download the data and make sure it makes sense.
- Zero out the randomization counts in the survey flow.
- Delete all of the fake responses.
- Upgrade your Qualtrics account with the code SDS290Fall2024.
- Activate your survey.
- Get the anonymous survey link!
Agenda
- Complete Block Design
Inappropriate Insects
Modern zoos try to reproduce natural habitats in their exhibits as much as possible. They try to use appropriate plants, but these plants can be infested with inappropriate insects. Cycads (plants that look vaguely like palms) can be infected with mealybugs, and the zoo wishes to test three treatments: 1) water, 2) horticultural oil, and 3) fungal spores in water. Nine infested cycads are taken to the testing area. Three branches are randomly selected from each tree, and 3 cm by 3 cm patches are marked on each branch. The number of mealybugs on the patch is counted. The three treatments then get randomly assigned to the three branches for each tree. After three days the mealybugs are counted again. The change in number of mealybugs is computed (\(before-after\)).
Inappropriate Insects
Draw the factor diagram.
MORE Design Terminology
- The two factors Trees and Treatment are crossed. This means that we collect data on all combinations of Trees and Treatment. A cell is a particular combination.
- Main effect refers to the effect for a single factor. For a block design there are two sets of main effects, for blocks and for treatments (e.g., Trees and Treatment).
- The two sets of main effects are called additive if each treatment adds or subtracts the same amount to the response variable for all blocks.
- A factor of interest is Treatment, which is directly related to the goal of the study. A nuisance factor is Trees which is included as a factor only because it is a source of variation and its inclusion will reduce the sum of squares for error (residuals).
Side-by-side dotplot
We add our blocking factor as color and also as group.
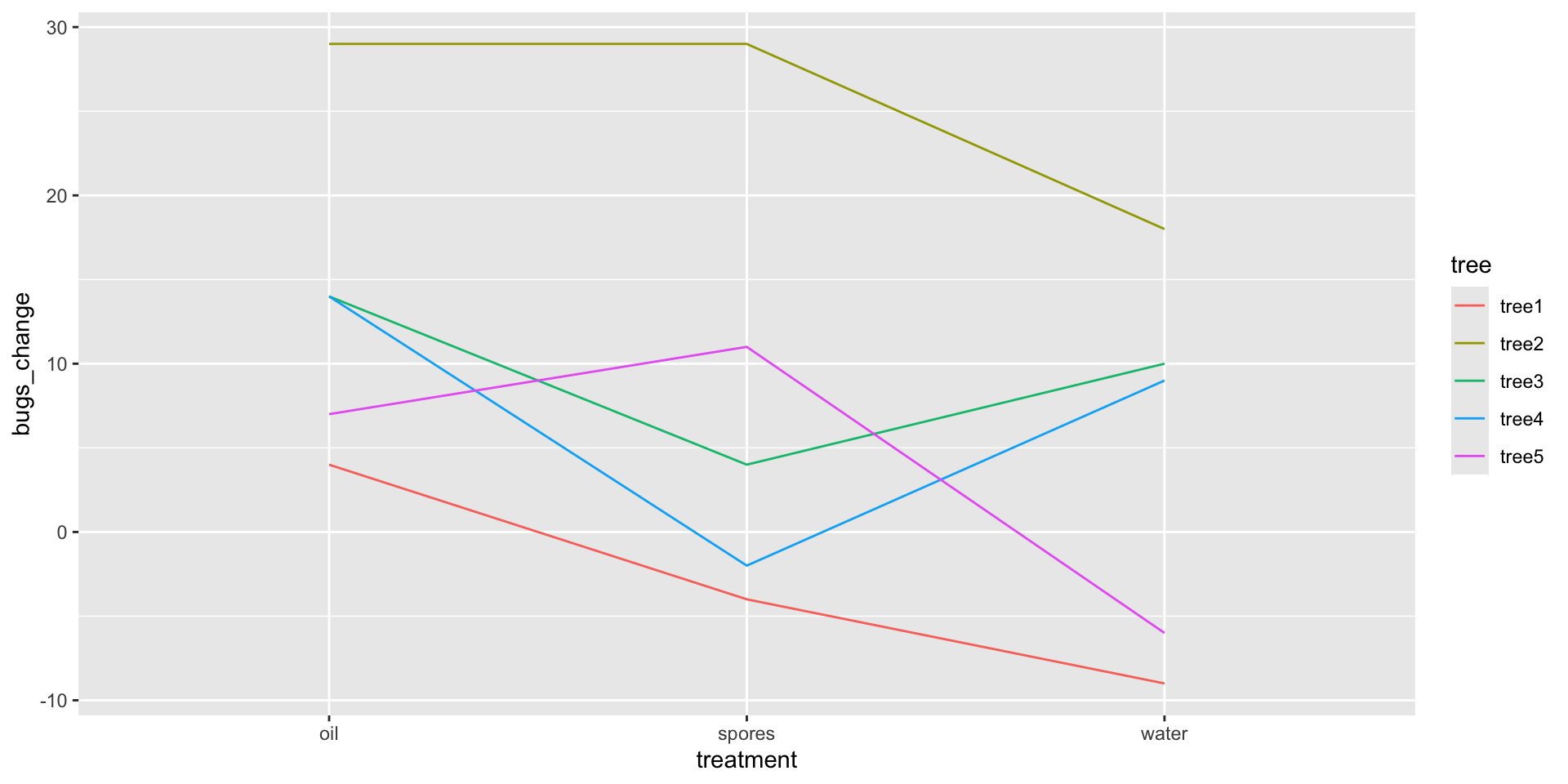
We can see that Tree 2’s infestation was very responsive to the treatments whereas Tree 1’s was not.
Formal ANOVA for Complete Block Design
\[{y}_{ij}={\mu}+{\tau}_{i}+{\beta}_{j}+{e}_{ij}\]
| Source | SS | df | MS | F |
|---|---|---|---|---|
| Treatment | \(\sum_{i=1}^{a}b(\bar{y}_{i.}-\bar{y}_{..})^{2}\) | \(a-1\) | \(\frac{{SS}_{T}}{{df}_{T}}\) | \(\frac{{MS}_{T}}{{MS}_{E}}\) |
| Blocks | \(\sum_{j=1}^{b}a(\bar{y}_{.j}-\bar{y}_{..})^{2}\) | \(b-1\) | \(\frac{{SS}_{B}}{{df}_{B}}\) | \(\frac{{MS}_{B}}{{MS}_{E}}\) |
| Error | \(\sum_{i=1}^{a}\sum_{j=1}^{b}({y}_{ij}-\bar{y}_{i.}-\bar{y}_{.j}+\bar{y}_{..})^{2}\) | \((a-1)(b-1)\) | \(\frac{{SS}_{E}}{{df}_{E}}\) |
Data Analysis Structure
Formal ANOVA
Analysis of Variance Table
Response: bugs_change
Df Sum Sq Mean Sq F value Pr(>F)
treatment 2 218.13 109.07 2.9963 0.106846
tree 4 1316.40 329.10 9.0412 0.004603 **
Residuals 8 291.20 36.40
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1Write a sentence interpreting the results of the F-test for treatment and a sentence interpreting the F-test for tree in the context of the problem.
Formal ANOVA
Analysis of Variance Table
Response: bugs_change
Df Sum Sq Mean Sq F value Pr(>F)
treatment 2 218.13 109.07 2.9963 0.106846
tree 4 1316.40 329.10 9.0412 0.004603 **
Residuals 8 291.20 36.40
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1There are no statistically significant differences in the reduction in mealy bugs between the three treatment conditions, \(F(2, 8) = 3.00\), \(p = .107\). There are significant differences in the reduction in mealy bugs across trees, however, \(F(4, 8) = 9.04\), \(p = .005\). That is, some trees improved more than other trees.
Informal Analysis Structure
How to check assumptions
C. Constant effects – think about whether it is reasonable.
A. Additive effects – check Anscombe block plots.
S. Same standard deviations – is the biggest SD less than two times as large as the smallest? check residual versus fitted plot: does the plot thicken?
I. Independent residuals – think about whether it is reasonable.
N. Normally distributed residuals – construct a histogram or normal probability plot of residuals.
Z. Zero mean residuals – construct a histogram or normal probability plot of residuals.
Anscombe Block Plots
- Scatterplots of two levels (e.g., level 1 vs. level 2) of the factor of interest.
- Used for
- exploring the data, and
- assessing the additivity (A) condition.
- If we compare a spore observation to an oil observation from the same tree, the only difference should be the difference in treatment effects.
- Thus, the line-of-best fit should have a slope of approximately 1.
Informal Analysis Structure
Anscombe Block Plots
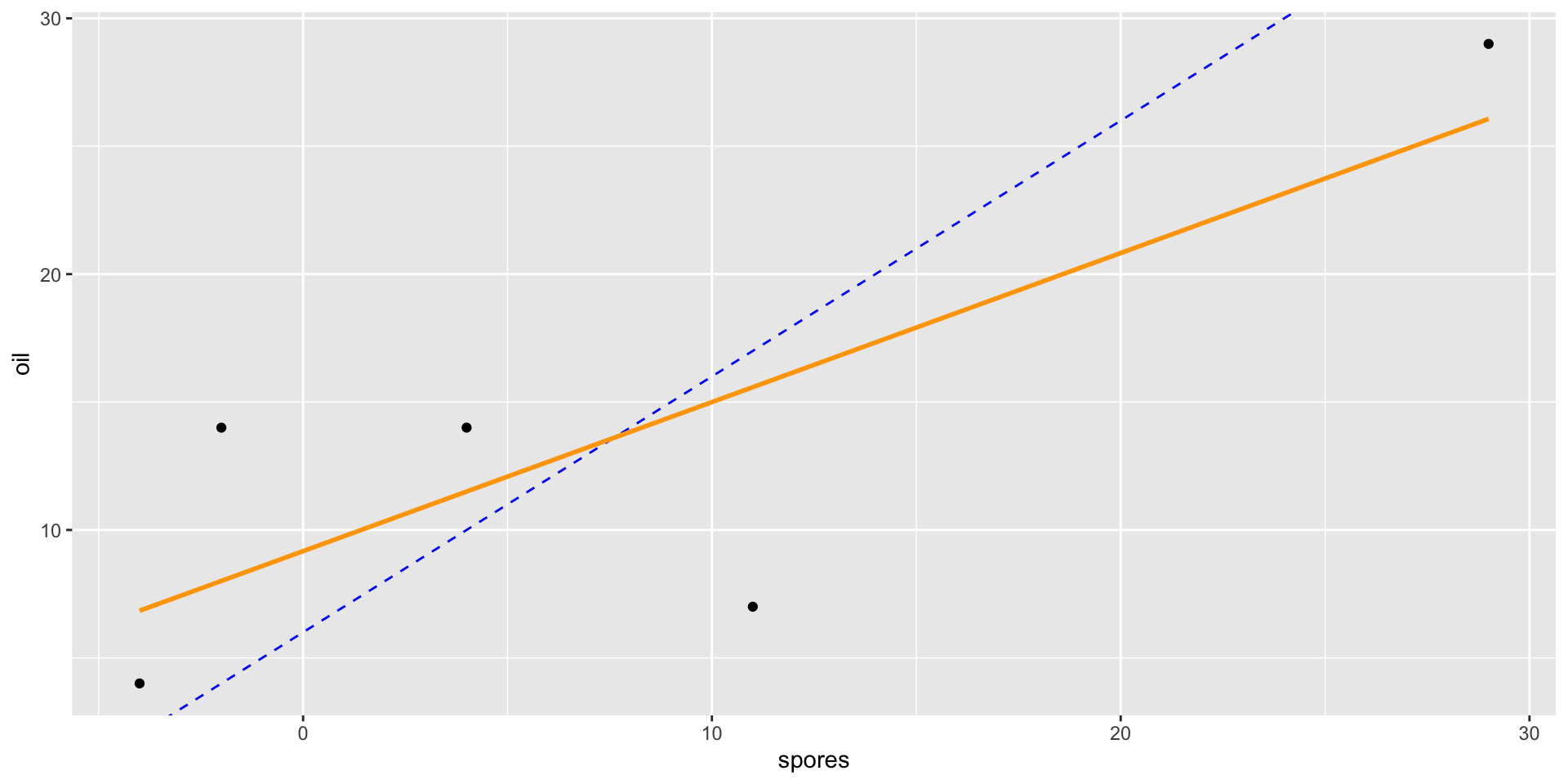
Anscombe Block Plots
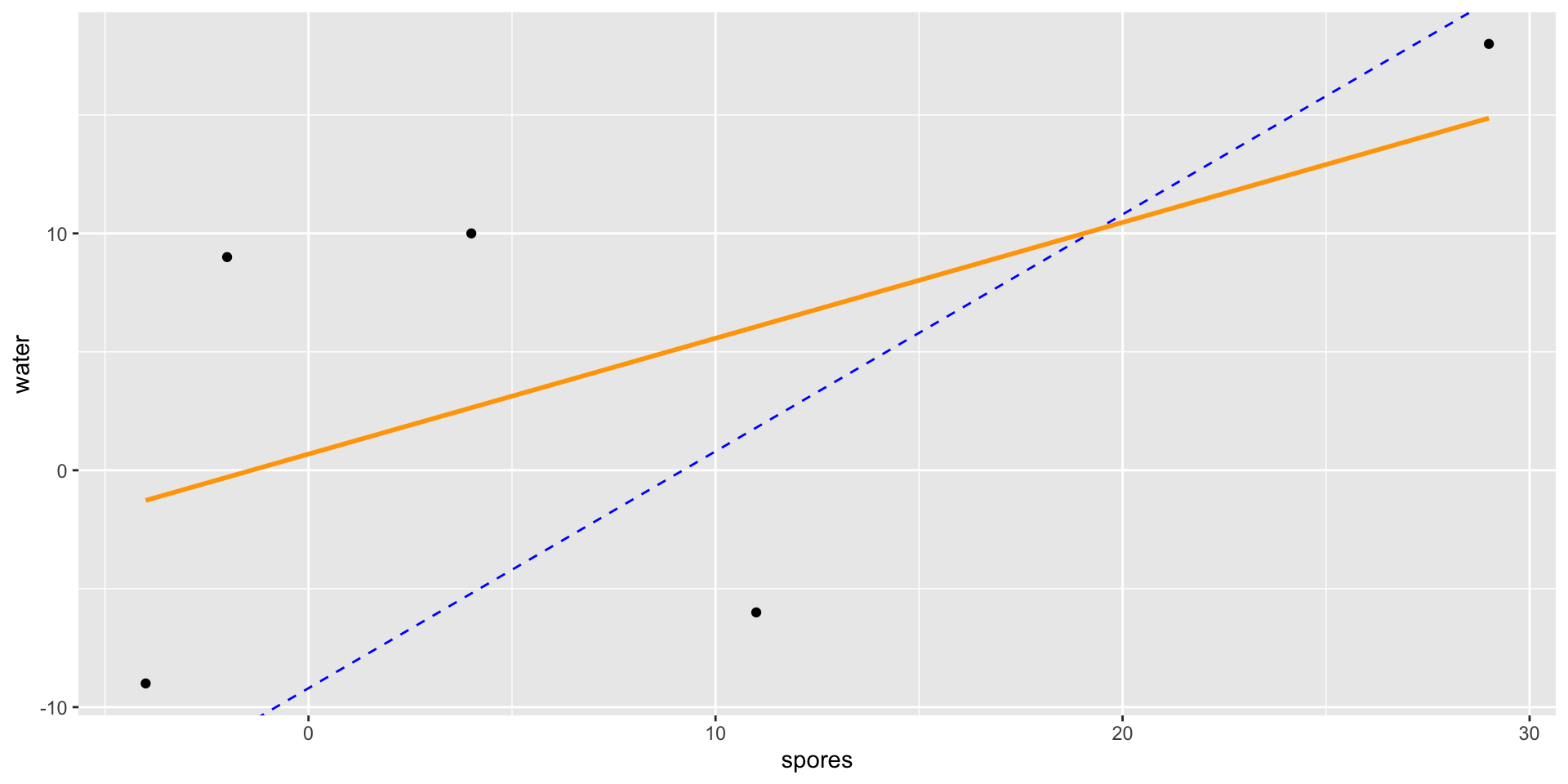
Anscombe Block Plots
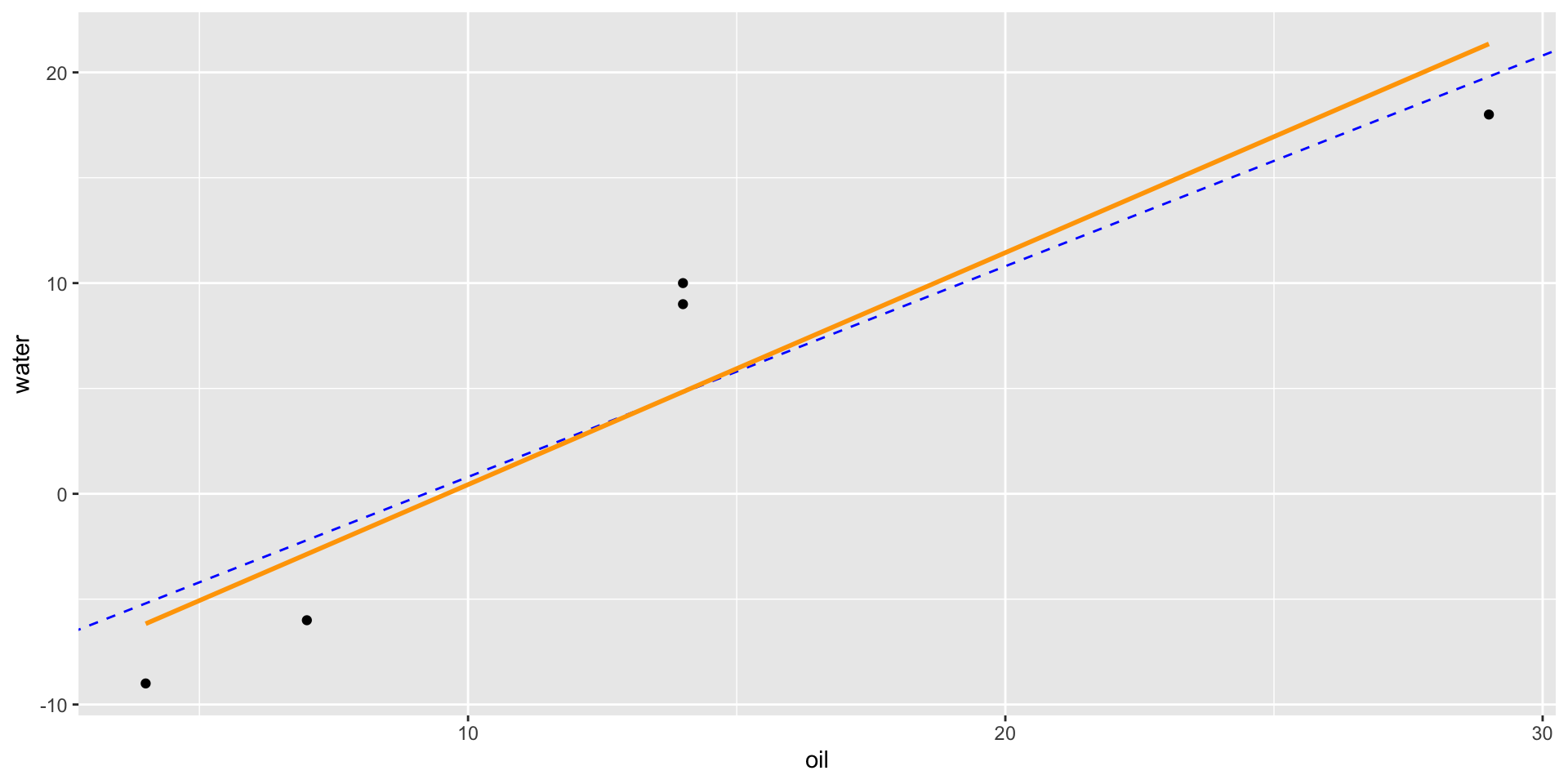
Assessing S Condition
# A tibble: 3 × 3
treatment m sd
<chr> <dbl> <dbl>
1 oil 13.6 9.66
2 spores 7.6 13.3
3 water 4.4 11.5 mealybugs %>%
group_by(treatment) %>%
summarize(m = mean(bugs_change),
sd = sd(bugs_change)) %>%
summarize(max(sd)/min(sd)) #calculating using min and max function# A tibble: 1 × 1
`max(sd)/min(sd)`
<dbl>
1 1.38Assessing S Condition
\[\hat{{y}}_{ij}={\mu}+{\tau}_{i}+{\beta}_{j}\]
- Where \(\hat{{y}}_{ij}\) are the fitted values, that is, everything but the ticket at the end of the assembly line.
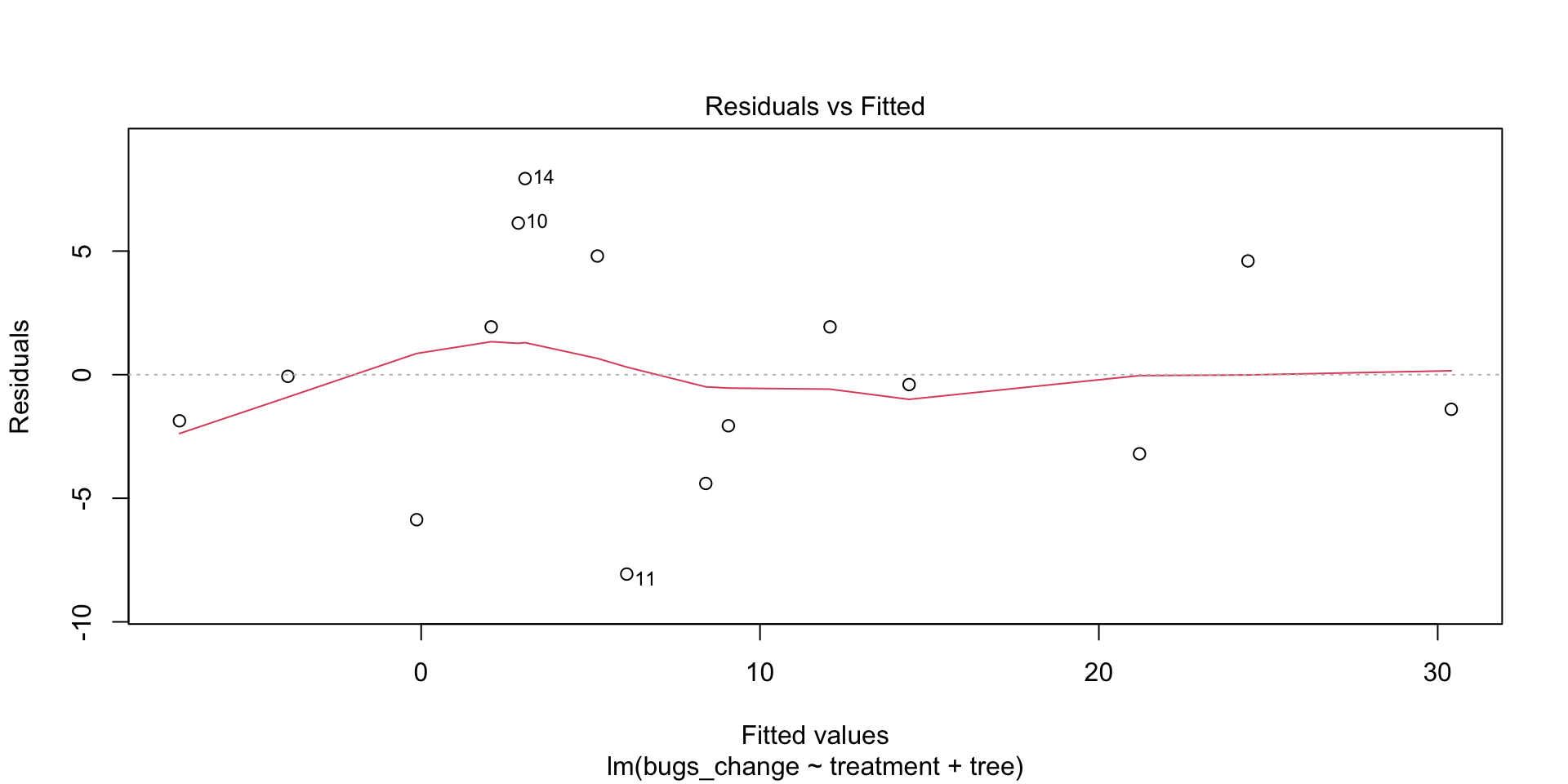
If the plot thickens, that is, has a patterning that looks like a funnel, then the S condition is not satisfied.
Assessing N Condition
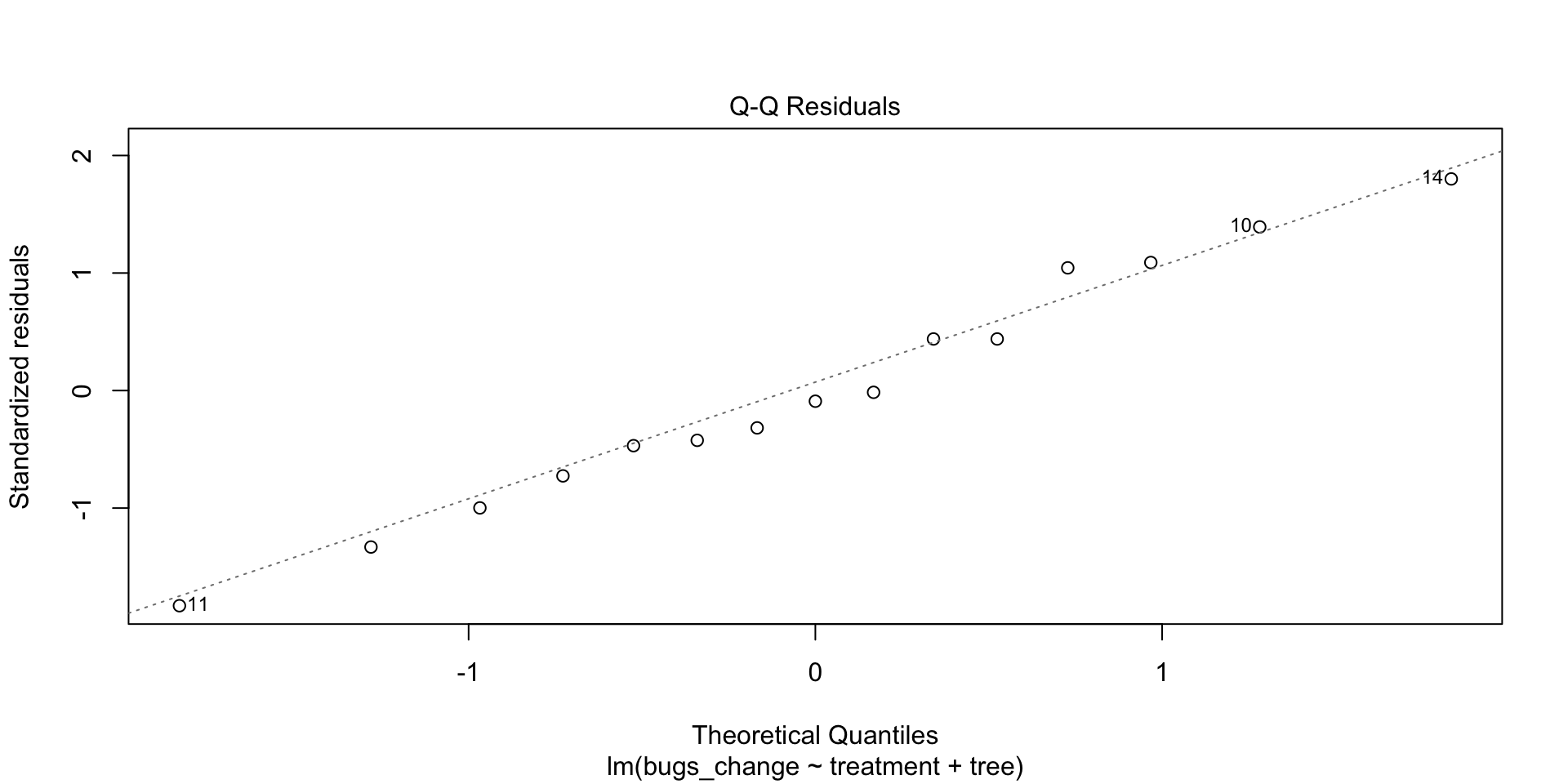
We’re looking for residuals to be on the line. If so, then we can say they are normally distributed.
Assessing the Z Condition
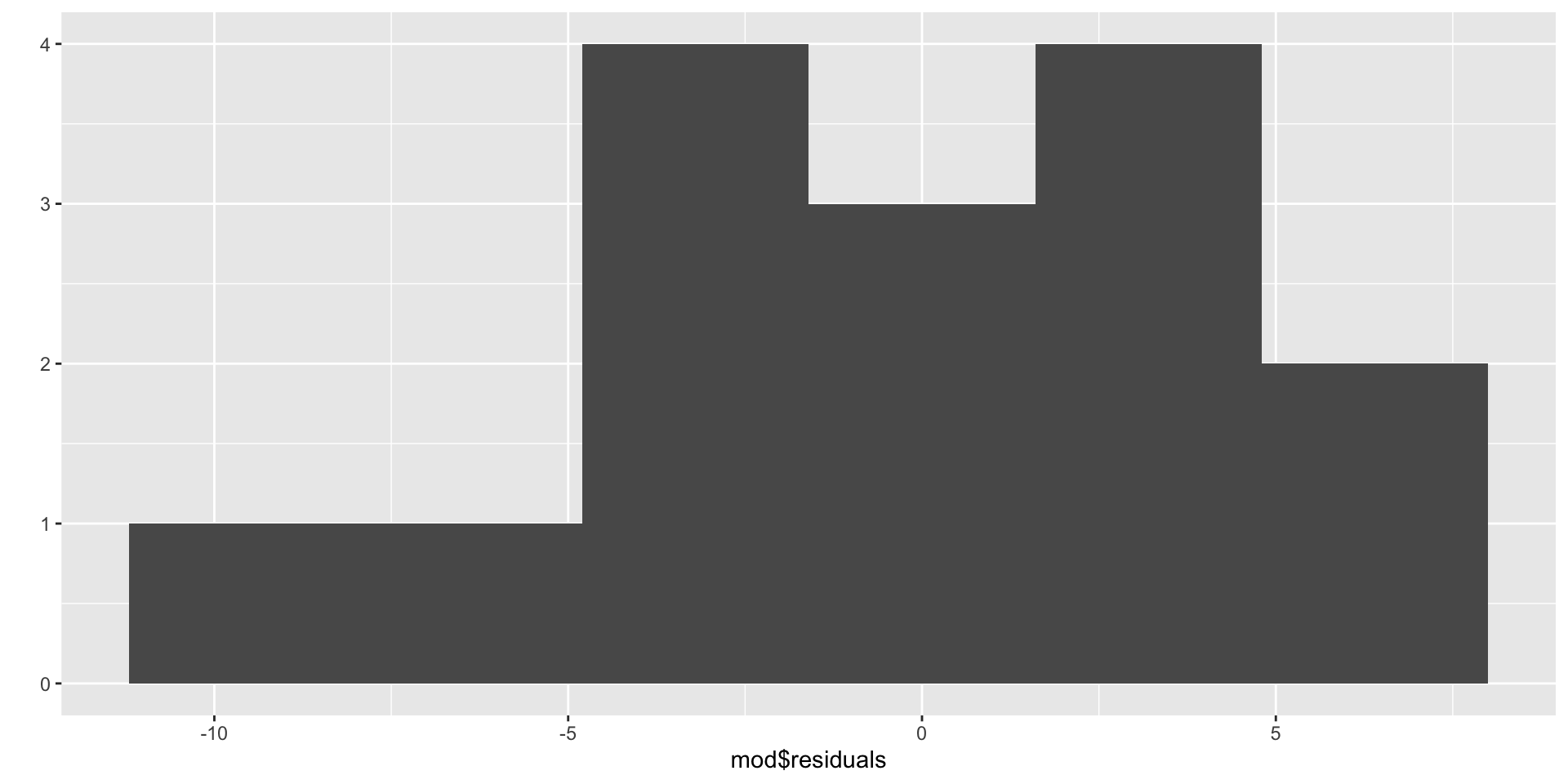
If the histogram centered at zero? Then the Z condition is satisfied.
Sleeping Shrews
Assess the CA-SINZ conditions for the SleepingShrews dataset from example 6.7b in your textbook.
Testing Condition for Sleeping Shrews Data
qplot(x = DSW, y = LSW, data = SleepingShrews_wide) +
geom_abline(intercept = 2, slope = 1, color = "blue", linetype = 2) +
geom_smooth(method = "lm", se = 0, color = "orange") #A condition
qplot(x = DSW, y = REM, data = SleepingShrews_wide) +
geom_abline(intercept = 2, slope = 1, color = "blue", linetype = 2) +
geom_smooth(method = "lm", se = 0, color = "orange") #A condition
qplot(x = LSW, y = REM, data = SleepingShrews_wide) +
geom_abline(intercept = -2, slope = 1, color = "blue", linetype = 2) +
geom_smooth(method = "lm", se = 0, color = "orange") #A condition
SleepingShrews %>%
group_by(Phase) %>%
summarize(m = mean(Rate),
sd = sd(Rate)) %>%
summarize(max(sd)/min(sd)) #S condition
mod <- lm(Rate ~ Phase + Shrew, data = SleepingShrews)
plot(mod, which = 1) #S condition
plot(mod, which = 2) #N condition
qplot(mod$residuals, bins = 6) #N and Z conditions